Finally, node levels of the sorted graph are mapped to the Cytoscape node attributes table. PSFC firstly performs depth first search traversal and removes backward edges from the graph, and performs the topological sorting on the resulting acyclic graph, after which the backward edges are restored. Recall that biological networks often contain feedback loops, which create cycles in graphs. ], to handle multiple input node containing graphs. We have modified the topological sort algorithm implemented in Java JGraphT library [ The aim of sorting is to assign levels to the nodes, to propagate the signal from lower to higher level nodes. Graph sorting is the first step before proceeding to signal flow calculation. Thus, it allows experimenting with the results obtained by various (existing and customizable) approaches within a single framework, and evaluating their ability to simulate real life situations. PSFC provides a variety of options for signal propagation, both used in already published signal flow algorithmsĦ, as well as in new ones. Herein we present Pathway Signal Flow Calculator (PSFC), a Cytoscape app for computation of pathway signal flow based on input data and pathway topology. 
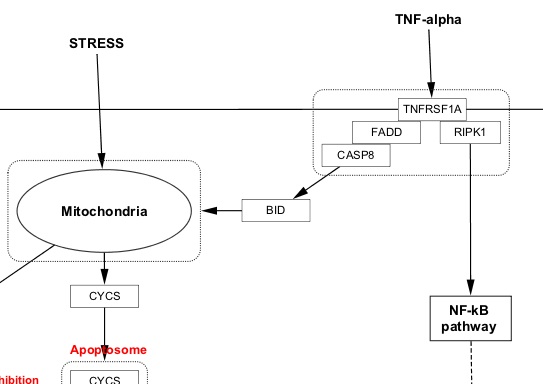
Cytoscape, on the other hand, is a powerful and flexible platform that, together with its diverse collection of available apps, provides a rich environment for parsing, visualization and analysis of networks Moreover, they often work with programming environment specific objects, and are not flexible for using biological pathways that appear in various formats. The above mentioned algorithms and tools are implemented using various programming and scripting languages, making their use and result comparison difficult in the common context. Other approaches apply specific rules to model flow or signal propagation through the pathway and evaluate the amount of the signal reaching the sink nodes One of the pioneering papers in this direction was the Pathway Impact Analysis algorithm, which combines GSEA with gene position in the networkģ. The major drawback of these widely used approaches is that they operate on gene sets involved in the pathway, but do not account for the pathway topology and ignore the interactions between the nodes.Ī number of techniques and tools have recently emerged, aimed at determining pathway activities based on topological information of pathways and gene expression/protein activity levels. Pathway involvement is typically analyzed by over-representation analysis (ORA)Ģ. First attempts to globally evaluate the pathway activity changes were performed in parallel with the appearance of high-throughput gene expression measurement experiments. Assessment of changes in pathway activity is of major interest for identification of processes involved in the formation of certain phenotypes (healthy and diseased states), and assessment of cell response to drugs and other stimuli. Thus, PSF can be an indicator of pathway activity state. Or perturbation, is the flux generated by propagation of the signal starting from input nodes, flowing through intermediate nodes in branches and accumulating at sink nodes. In contrast to protein-protein interaction networks, biomolecular pathways have directionality, input nodes, intermediate nodes and branches, and output or sink nodes. Pathways can be represented as graphs, with nodes as biological entities (proteins, other biomolecules, chemical compounds, other pathways), and edges as physical or regulatory interactions between them. a ligand binding to a receptor) and result in realization of certain target processes (e.g. Cell signaling pathways are sets of directed interactions between biological molecules, that are initiated by a particular signal (e.g. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
